Monday at Darwin’s, March Madness Day 5
ø
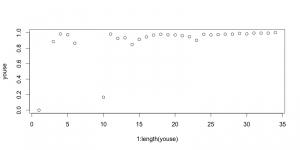
The weekend was not super-productive, although I did put in some time on both days. I was pretty exhausted on Saturday after spending the morning volunteering at the museum, and spent some time napping at Pierre and Nicole’s while Kati babysat Alexandre. In any case I managed to spend several hours over the course of the rest of the weekend tidying up the subsampled plot so I’m now satisfied with the way it looks:
This done, I moved on to tackling the next task, which is to try and answer Andy’s (legitimate) question about which characters are actually responsible for changes in morphospace occupancy. To accomplish this, I wrote code that compares the taxa in the alpha volumes of adjacent time bins and finds the taxa in the new, expanded volume of the younger time bin (i.e. those taxa that fall outside of the volume of the older time bin). The code then compares the character states of those taxa with the taxa in the older time bin and identifies which character states are new. Those character states are responsible for the expansion of morphospace volume (at least in three dimensions).
This new chunk of code returns lists of character states for each time bin, which is of course a little obscure, being a list of numbers, and requires a bit of look-up work. Here, then, is a summary of the character states that are responsible for adding alpha volume in each time bin. (This took a bit of time, too, because I decided it was too laborious to look up each of the character descriptions, so I wrote a script to call up the appropriate line of description from the text file containing the character and state descriptions).
- Late Cretaceous: 68 new states outside the Early Cretaceous 3D-morphospace alpha volume (with alpha=0.11). A lot of states to look at here. Yikes. New valve outline shapes, aspect ratios, undulations, torsion, and curvature. Apex shape, heterovalvy, topographic folds, apex topography, various other characters of the apex, central depression, ornament at rim, asymmetric mantle, warts, brim, mantle pores, surface texture, central area ornament, spinules, collar. Importantly: the first sternum. Pore arrangement and size, other pore features, bullulae, pseudoseptae & ribs, apical fields, labiate processes, some sternum characters.
- Paleocene: 53 new states outside the Late Cretaceous volume. Again, a lot. Ovate valve outline, depresse aspect ratio, frustule curvature, apiculate apices, heterovalvy, valves without a mantle, crimped mantle edge, mantles without pores or with simple perforations, more costa types, rays & associated characters, non-perpendicular pore arrangement, oval and rectangular pores, alveoli, porelli, pseudonoduli, ocelli, apical pore fields in rows, wide as well as eccentric sternum. Importantly, the raphe, in various places, singly and split, with and without canal, keel, and fibulae.
- Eocene: 17 new states outside the Paleocene volume. Rhombic valve outline, curved central elevation, more costae, central area with scattered pores, central spines or tubercles, curved and spiral rows of pores, quadrate and slit-like pores, apical and annular pseudosepta, ocelluli, sigmoidal raphes, and laterally-opening raphes.
- Oligocene: 2 new states outside the Eocene volume. Naviculoid non-sternum central area, ring of tubular spines in central area. (Whoa, random—I guess these are the only things Pseudorutilaria has that weren’t there pre-Oligocene; it’s just its odd combination of all the other characters that makes it stand out. I guess.).
- Miocene: 16 new states outside the Oligocene volume. Panduriform valve outline, curved frustule, patchy pores on mantle, ridges and grooves on valve face, biseriate pores, specialized openings, quite a few invalid states*, labiate processes along sternum, sinuous raphe, straight, deflected, and bent terminal raphe fissures.
- Plio-Pleistocene: 4 new states outside the Late Cretaceous volume. Anguste late aspect ratio, sternum widening at poles, raphe around entire valve circumference, and one invalid state*.
*It was pretty shocking to see how many invalid states there were. These are, for example, a state 3 when there are only states 0, 1, and 2, or in one case a state “j”. These are obviously typos. Clearly, they’re more likely to be caught by looking at taxa in the fringes of the morphospace and by looking at characters not found in the other taxa, but it’s still kind of freak-out-ish to think how many typos and other mistakes are lurking in my dataset. Alas, I simply don’t have the time to fix these anymore. Going through the whole dataset (even the culled dataset contains 14,000 entries) is just not feasible. End of story. The mistakes have to stay in there.
Is there anything to be learned from the characters responsible for the space expansion? Nothing jumps out at me. The shift from Early to Late Cretaceous is uninteresting to me, just because there are so few taxa in the Early Cretaceous that I doubt it represents much of anything real; anyhow, the characters are mostly related to seeing pennate diatoms. In the Paleocene… I don’t know. Just seems like a random jumble of characters. Same for the other time bins really. What a pile of crap shit poop. My guess is that the morphospace is just basically meaningless, so the characters responsible for these volume gains are essentially random.
So what remains to be done now? I have not, as promised in my to-dos for the week, spent the requisite time on my thesis document, writing, so that should probably be my next task for today, if only briefly, so that I don’t lose touch completely again. Got a couple of paragraphs on the morphological vs. molecular distances written up. Slow like treacle, but it’s coming along.