Time to Put the Cards on the Table
ø
After a weekend that was less productive than I would have hoped, I finally bit the bullet and made a start on laying out my index cards and building the structure of the papers. It’s been quite helpful so far—forcing myself to determine what conclusions I can actually make with the data and plots I have makes it clear that those are the questions I must ask at the outset, and build introductions to support.
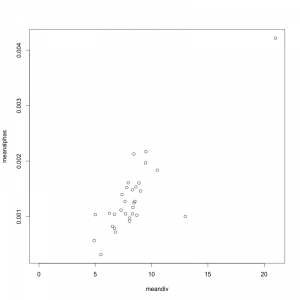
A big sticking point for me right now is the first paper. With the exception of the side-track about morphology and phylogeny, this paper is basically about “how to build a morphospace”, which isn’t really an interesting finding so much as a long methods section. The one thing I realized in doing it, and that Andy found quite interesting too, is that the choice of data culling criteria can have a pretty substantial effect on what you see, and it’s not a choice that has (to my knowledge) been addressed explicitly in prior morphospace studies. But for that to be a useful finding of the paper, I will need to actually run some analyses to show how different choices affect the outcome. Setting up the choices shouldn’t be too hard—I should think starting with the full data set as collected and then executing random (bootstrap) replicates for progressively smaller subsets of the original data is the way to go—but I’m not sure what metric I should use to show these effects. Mean pairwise distance through time? This is one of those metrics that is used a lot, but then I need to front-load a whole lot of explanation about the through-time stuff (linking the morphospace to Neptune, etc.) that I was hoping to save for the second paper, where I think this belongs.
Anyway, these are the sorts of questions I’m dealing with. I really had hoped that I was all done with analyses by this point, but I’m not sure how well the first paper will stand up on its own without a little bit of additional work. It’s really not a huge deal—it shouldn’t take more than a day to code up once I’ve decided on a metric—but choosing an output variable that captures what I’m trying to say and works with the logic and construction of the two papers is a bit of a challenge.
Here is what the poker table looks like, by the way:
I did also take two closer-up views of the layout for the first paper and the second paper, just in case disaster strikes in the form of wind, fire, loss, etc. You can’t be too careful at this stage.